Plasmodium falciparum Kelch 13-propeller gene mutation update in Nigeria – a systematic review
DOI:
https://doi.org/10.60787/tnhj.v23i2.670Keywords:
K13-propeller gene polymorphism, Artemisinin resistance, Malaria Risk, Plasmodium falciparum, NigeriaAbstract
Background: Mutations in Plasmodium falciparum Kelch 13 propeller gene has been associated with artemisinin resistance. This review is a synthesis of evidence from research on Plasmodium falciparum Kelch 13 (Pfk13) propeller gene mutations conducted in Nigeria from 2011 to 2023 to determine the extent of spread of Pfk13 mutations in different states of Nigeria.
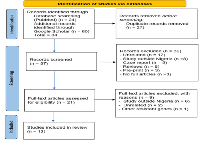
Methods: An electronic search of studies from 2011 to date was done in Medline via PubMed, database and Google Scholar using Mesh terms and Boolean operators. PRISMA guidelines were used to guide the screening of articles for inclusion. Data extraction form was developed in Excel 2016 and used for data extraction.
Results: A total of 84 articles were retrieved but only 12 were eligible for inclusion. Pfk13 gene mutations studies have been conducted in 11 out of 36 (30.6%) states of Nigeria. The total number of independent Pfk13 mutations identified across the different states was 44, 32 (72.7%) mutations were novels. None of the reported mutations in this review was among the validated k13 mutations associated with increased artemisinin resistance.
Conclusion: Number of Pfk13 gene mutations studies in Nigeria was scanty. States in the North-east, North-west, and South-south geopolitical regions of Nigeria have the poorest coverage. Pfk13 gene mutations including novel ones were identified, however, they were all non-validated k13 mutations. There is need to carry out further studies (in vitro and in vivo) to ascertain the role of these novel mutations in emergence and spread of ART resistance in Nigeria.
Downloads
References
WHO. Malaria: Artemisinin resistance [Internet]. 2023 [cited 2023 Mar 11]. Available from: https://www.who.int/news-room/questions-and-answers/item/artemisinin-resistance
Iwuafor AA, Ogban GI, Emanghe UE, Erengwa PC, Offiong AB, Nsor GE, et al. Artemisinin drug resistance and monitoring: a narrative review. African J Clin Exp Microbiol. 2023;24(2):112–9.
Ariey F, Witkowski B, Amaratunga C. A molecular marker of artemisinin-resistant Plasmodium falciparum malaria. Nature. 2014;505:50–5.
Miotto O, Amato R, Ashley EA, MacInnis B, Almagro-Garcia J, Amaratunga C, et al. Genetic architecture of artemisinin-resistant Plasmodium falciparum. Nat Genet. 2015 Mar;47(3):226–34.
Adams J, Kelso R, Cooley L. The kelch repeat superfamily of proteins: propellers of cell function. Trends Cell Biol. 2000;10:17–24.
Straimer J, Gnädig NF, Witkowski B, Amaratunga C, Duru V, Ramadani AP, et al. Drug resistance. K13-propeller mutations confer artemisinin resistance in Plasmodium falciparum clinical isolates. Science (80-). 2015;347(6220):428–31.
WWARN K13 Genotype-Phenotype Study Group. Association of mutations in the Plasmodium falciparum Kelch13 gene (Pf3D7_1343700) with parasite clearance rates after artemisinin-based treatments-a WWARN individual patient data meta_analysis. BMC Med. 2019;17(1):1.
Ng CL, Fidock DA. Plasmodium falciparum in vitro drug resistance selections and gene editing. Methods Mol Biol. 2019;2013:123–40.
Rosenthal MR, Ng CL. Plasmodium falciparum artemisinin resistance: The effect of heme, protein damage, and parasite cell stress response. ACS Infect Dis. 2020;6:1599–1614.
Talman AM, Clain J, Duval R, Menard R, Ariey F. Artemisinin bioactivity and resistance in malariaparasites. Trends Parasitol. 2019;35:953–963.
Nsanzabana C, Ariey F, Beck HP. Molecular assays for antimalarial drug resistance surveillance: a target product profile. PLoS One. 2018;13:e0204347.
Talisuna AO, Karema C, Ogutu B, Juma E, Logedi J, Nyandigisi A, et al. Mitigating the threat of artemisinin resistance in Africa: improvement of drug-resistance surveillance and response systems. Lancet Infect Dis. 2012;12:888–96.
Uwimana A, Legrand E, Stokes B, Ndikumana J, Warsame M, Umulisa N.Emergence and clonal expansion of in vitro artemisinin-resistant Plasmodium falciparum kelch13 R561H mutant parasites in Rwanda. Nat Med. 2020;26:1602–1608.
L’Episcopia M, Bartoli T, Corpolongo A. Artemisinin resistance surveillance in African Plasmodium falciparum isolates from imported malaria cases to Italy. J Travel Med. 2021;28(5):taaa231.
Balikagala B, Fukuda N, Ikeda M, Katuro O, Tachibana SI, Yamauchi M. Evidence of artemisinin-resistant malaria in Africa. N Engl J Med. 2021;385:1163–1171.
WHO. Methods for surveillance of antimalarial drug efficacy. Geneva: World Health Organisation; 2009.
Kamau E, Campino S, Amenga-Etego L, Drury E, Ishengoma D, Johnson K, et al. K13-propeller polymorphisms in Plasmodium falciparum parasitesfrom sub-Saharan Africa. J Infect Dis. 2015;211(8):1352–5.
Oboh M, Ndiaye D, Antony H, Badiane A, Singh U, Ali N, et al. Status of artemisinin resistance in malaria parasite Plasmodium falciparum from molecular analyses of the Kelch13 gene in Southwestern Nigeria. Biomed Res Int [Internet]. 2018 [cited 2023 Jan 21]; Available from: https://www.hindawi.com/journals/bmri/2018/2305062/
Igbasi U, Oyibo W, Omilabu S, Quan H, Chen S-BB, Shen H-MM, et al. Kelch 13 propeller gene polymorphism among Plasmodium falciparum isolates in Lagos, Nigeria: Molecular Epidemiologic Study. 2019 Aug 1 [cited 2023 Jan 21];24(8):1011–7. Available from: https://onlinelibrary.wiley.com/doi/abs/10.1111/tmi.13273
Idowu AO, Oyibo WA, Bhattacharyya S, Khubbar M, Mendie UE, Bumah V V., et al. Rare mutations in Pfmdr1 gene of Plasmodium falciparum detected in clinical isolates from patients treated with anti-malarial drug in Nigeria. Malar J. 2019 Sep 18;18(1).
Olasehinde G, Diji-Geske R, Fadina I, Arogundade D, Darby P, Adeleke A, et al. Epidemiology of Plasmodium falciparum infection and drug resistance markers in Ota Area, Southwestern Nigeria. Infect Drug Resist. 2019;12:1941–9.
Tola M, Ajibola O, Idowu E, Omidiji O, Awolola S, Amambua-Ngwa A. Molecular detection of drug resistant polymorphisms in Plasmodium falciparum isolates from Southwest, Nigeria. BMC Res Notes. 2020;13:497.
Abubakar U, Adam R, Mukhtar M, Muhammad A, Yahuza A, Ibrahim S. Identification of mutations in antimalarial resistance gene kelch13 from Plasmodium falciparum isolates in Kano, Nigeria. Trop Med Infect Dis. 2020;5(85).
Ikegbunam M, Ojo JA, Kokou K, Morikwe U, Nworu C, Uba C, et al. Absence of Plasmodium falciparum artemisinin resistance gene mutations eleven years after the adoption of artemisinin-based combination therapy in Nigeria. Malar J. 2021 Dec 1;20(1).
Muhammad I, Sale P, Midala A. Absence of Biomakers of Resistance in K13 Propeller Gene of Plasmodium falciparum from Gombe LGA of Gombe State, Nigeria. Advances. 2022;3(1):25–33.
Ajogbasile F, Oluniyi P, Kayode A, Akano K, Adegboyega B, Philip C, et al. Molecular profiling of the artemisinin resistance Kelch 13 gene in Plasmodium falciparum from Nigeria. PLoS One. 2022;17(2):e0264548.
Afolabi O, Oluwafemi O, Diseases MO-J of P, 2022 U. Pfmdr 1 and kelch 13 genes distribution among children that are 5 years and below in Akure, Nigeria. J Parasit Dis. 2022;47(1):59–67.
Adulugba O, Amali O, Manyi M, Ikpa T, Obisike V. Genetic Diversity and Molecular Surveillance of Antimalarial Drug Resistance of Plasmodium falciparum among Hospitals Patients in Benue State Nigeria. Microbiol Res J Int. 2022;32(1):1–10.
Ocan M, Akena D, Nsobya S, Kamya MR, Senono R, Kinengyere AA, et al. K13-propeller gene polymorphisms in Plasmodium falciparum parasite population in malaria affected countries: a systematic review of prevalence and risk factors. Malar J. 2019 Mar;18(1):60.
World Malaria Report 2021: An in-depth update on global and regional malaria data and trends. [Internet]. Geneva: World Health Organisation; 2021. Available from: https://www.who.int/teams/global-malaria-programme/reports/world-malaria-report-2021
WHO. Malaria threats map. Geneva: World Health Organization [Internet]. 2021 [cited 2021 Dec 30]. Available from: https://apps.who.int/malaria/maps/threats/
Asua V, Vinden J, Conrad MD, Legac J, Kigozi SP, Kamya MR, et al. Changing Molecular Markers of Antimalarial Drug Sensitivity across Uganda. Antimicrob Agents Chemother. 2019 Mar;63(3).
Straimer J, Gandhi P, Renner KC, Schmitt EK. High Prevalence of Plasmodium falciparum K13 Mutations in Rwanda Is Associated With Slow Parasite Clearance After Treatment With Artemether-Lumefantrine. J Infect Dis. 2022 Apr;225(8):1411–4.
Conrad M, Bigira V, Kapisi J, Muhindo M, Kamya M, Havlir D. Polymorphisms in K13 and falcipain-2 associated with artemisinin resistance are not prevalent in Plasmodium falciparum isolated from Ugandan Children. PLoS One. 2014;9:e105690.
Boussaroque A, Fall B, Madamet M, Wade KA, Fall M, Nakoulima A, et al. Prevalence of anti-malarial resistance genes in Dakar, Senegal from 2013 to 2014. Malar J. 2016 Jul;15(1):347.
Ashley E, Dhorda M, Fairhurst R, Amaratunga C, Lim P, Suon S. Spread of artemisinin resistance in Plasmodium falciparum malaria. N Engl J Meg. 2014;371:411–23.
Talundzic E, Ndiaye Y, Deme A, Olsen C, Patel D, Biliya S. Molecular epidemiology of Plasmodium falciparum kelch13 mutations in Senegal determined by using targeted amplicon deep sequencing. Antimicrob Agents Chemother. 2017;61:2116.
Boussaroque A, Fall B, Madamet M, Camara C, Benoit N, Fall M, et al. Emergence of mutations in the K13 propeller gene of Plasmodium falciparum isolates from Dakar, Senegal, in 2013-2014. Antimicrob Agents Chemother. 2016 Jan 1;60(1):624–7.
Yang C, Zhang H, Zhou R, Qian D, Liu Y, ... YZ-B infectious, et al. Polymorphisms of Plasmodium falciparum k13-propeller gene among migrant workers returning to Henan Province, China from Africa. BMC Infect Dis. 2017;17(1):560.
Gaye A, Sy M, Ndiaye T, Siddle KJ, Park DJ, Deme AB, et al. Amplicon deep sequencing of kelch13 in Plasmodium falciparum isolates from Senegal. Malar J. 2020;19(1).
Downloads
Published
Versions
- 2023-07-20 (2)
- 2023-07-10 (1)
Issue
Section
License
Copyright (c) 2023 Journal and Publisher

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
The Journal is owned, published and copyrighted by the Nigerian Medical Association, River state Branch. The copyright of papers published are vested in the journal and the publisher. In line with our open access policy and the Creative Commons Attribution License policy authors are allowed to share their work with an acknowledgement of the work's authorship and initial publication in this journal.
This is an open access journal which means that all content is freely available without charge to the user or his/her institution. Users are allowed to read, download, copy, distribute, print, search, or link to the full texts of the articles in this journal without asking prior permission from the publisher or the author.
The use of general descriptive names, trade names, trademarks, and so forth in this publication, even if not specifically identified, does not imply that these names are not protected by the relevant laws and regulations. While the advice and information in this journal are believed to be true and accurate on the date of its going to press, neither the authors, the editors, nor the publisher can accept any legal responsibility for any errors or omissions that may be made. The publisher makes no warranty, express or implied, with respect to the material contained herein.
TNHJ also supports open access archiving of articles published in the journal after three months of publication. Authors are permitted and encouraged to post their work online (e.g, in institutional repositories or on their website) within the stated period, as it can lead to productive exchanges, as well as earlier and greater citation of published work (See The Effect of Open Access). All requests for permission for open access archiving outside this period should be sent to the editor via email to editor@tnhjph.com.